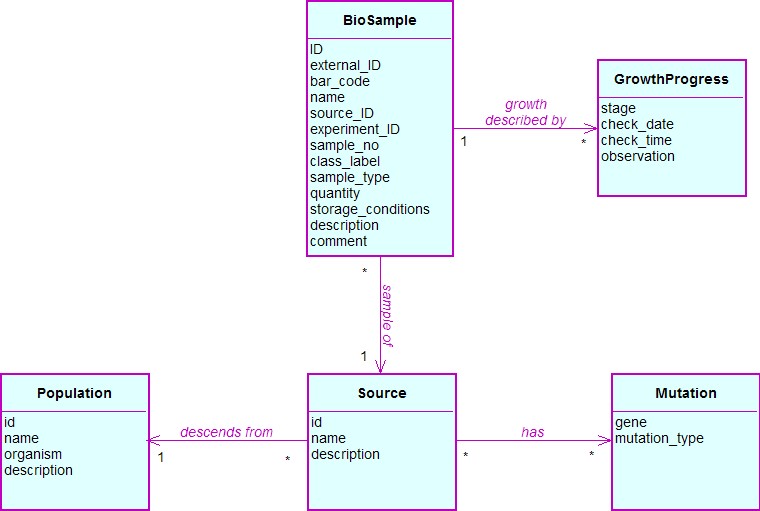
Biological sample
Each metabolomics study concentrates on a specific biological source. An instance of the Source class represents an individual organism (e.g. a specific patient in case of clinical studies) or a genetically defined strain of a particular organism (e.g. a specific gene-knockout strain of S. cerevisiae). In this model we focus only on the latter case, where each Source instance is linked to instances of the Mutation class denoting the genes that were knocked out/in from a given biological source. Large-scale genomics studies investigate whole populations of related biological sources rather than a single source. For example, there is sometimes a mutant for each gene in functional genomics studies using a model organism such as S. cerevisiae. Instances of the Population class are used to describe a studied group of related biological sources. Each Source is used to produce a biological sample (e.g. metabolomic footprint of a given S. cerevisiae strain), which is represented using the BioSample class.